Visit my

GitHub page
and

Bitbucket page.
Multilayer Networks Library
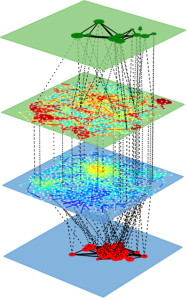
Main features
- Pure Python
- Can handle general multilayer networks
- Multilayer networks and multiplex networks (with automatically generated lazy-evaluation coupling edges)
- Functionality: Analysis, transformations, reading and writing networks, network models etc.
- Visualization (using Matplotlib or D3)
- Integration with NetworkX for monoplex network analysis
Get it from Github.
EDENetworks
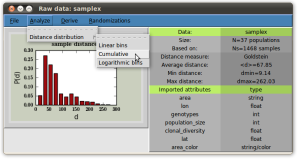
EDENetworks is an easy to use tool for analysing genetic networks. It allows researchers to use the tools from complex networks science to analyse genetic data using a user friendly graphical interface. You can study both individual level or population level networks by giving either a genetic marker file or a distance matrix as an input.
Visit the EDENetworks website.
